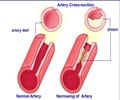
Explore the intriguing question of whether fragmented coding regions can fold into functional proteins.
Reassessing the exon-foldon correspondence using frustration analysis
Go to source). The researchers focused on the extensive relationship between exons in protein structures and the evolution of protein foldability. They highlighted the significance of exons, the parts of the gene that code for proteins, and introns, the silent regions discarded during gene translation into proteins.
‘#Foldableproteins are biomolecules composed of amino acids that can adopt specific three-dimensional structures, often spontaneously or with the help of chaperones. These structures are critical for their biological functions’





Investigating Exon Boundary Conservation with Energy Landscape Theory
“Using the extensive genomic exon-intron organization and protein sequence data now available, we explored exon boundary conservation and assessed its behavior using energy landscape theoretic measurements,” said Wolynes, the D.R. Bullard-Welch Foundation Professor of Science, professor of chemistry, biosciences, physics and astronomy and co-director of the Center for Theoretical Biological Physics (CTBP).When genes in pieces were discovered in the 1970s, it was immediately proposed that by breaking up the sequence, this structure helped build foldable proteins. When researchers looked at this again in the 1990s, the existing data was equivocal, Wolynes said.
The team has now assessed exons as potential protein folding modules across 38 abundant and conserved protein families. Over generations, exons can shuffle randomly along the genome, leading to significant changes in genes and the creation of new proteins. The findings indicated deviations in the exon size distribution from exponential decay, suggesting there was evolutionary selection.
“Protein folding and evolution are closely linked phenomena,” said Ezequiel Galpern, a postdoctoral researcher at the University of Buenos Aires.
Natural proteins are linear chains of amino acids that typically fold into compact three-dimensional structures to perform biological functions. The specific sequence of amino acids dictates the final 3D structure. Therefore, the idea that exons translate into independently folded protein regions, or foldons, is very attractive.
Advertisement
The study found a correlation between protein folding and evolution in certain globular protein families. Protein folding involves amino acid chains folding in space to perform biological functions within relevant timescales. This correlation is a fundamental concept in protein science, assessed using genomic data and energy functions.
Advertisement
Reference:
- Reassessing the exon-foldon correspondence using frustration analysis - (https://www.pnas.org/doi/10.1073/pnas.2400151121)
Source-Eurekalert