Deep-learning computer network could help to automate biopsy slides for breast cancer, reveals study.
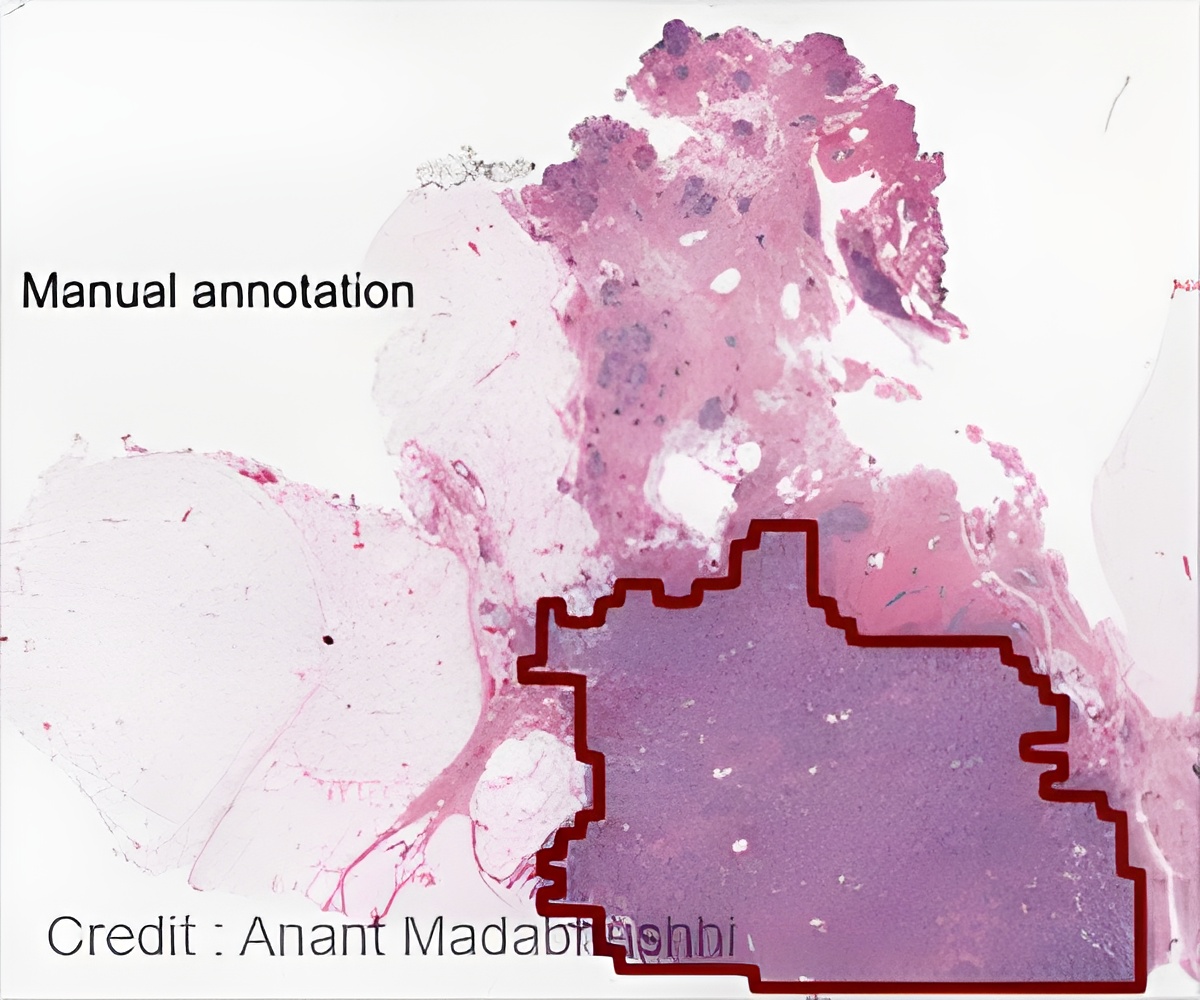
Compared to the analyses of four pathologists, the machine was more consistent and accurate, in many cases improving on their delineations.
In a field where time and accuracy can be critical to a patient's long-term prognosis, the study is a step toward automating part of biopsy analysis and improving the efficiency of the process, the researchers say.
Currently, cancer is present in one in 10 biopsies ordered by physicians, but all must be analyzed by pathologists to identify the extent and volume of the disease, determine if it has spread and whether the patient has an aggressive or indolent cancer and needs chemotherapy or a less drastic treatment.
Last month, the U.S. Food and Drug Administration approved software that allows pathologists to review biopsy slides digitally to make diagnosis, rather than viewing the tissue under a microscope.
"If the network can tell which patients have cancer and which do not, this technology can serve as triage for the pathologist, freeing their time to concentrate on the cancer patients," said Anant Madabushi, F. Alex Nason professor II of biomedical engineering at Case Western Reserve and co-author of the study detailing the network approach, published in Scientific Reports.
Advertisement
To train the deep-learning network, the researchers downloaded 400 biopsy images from multiple hospitals. Each slide was approximately 50,000 x 50,000 pixels. The computer navigated through or rectified the inconsistencies of different scanners, staining processes and protocols used by each site, to identify features in cancer versus the rest of the tissue.
The researchers then presented the network with 200 images from The Cancer Genome Atlas and University Hospitals Cleveland Medical Center. The network scored 100 percent on determining the presence or absence of cancer on whole slides and nearly as high per pixel.
Advertisement
Network training took about two weeks, and identifying the presence and exact location of cancer in the 200 slides took about 20 to 25 minutes each.
That was done two years ago. Madabhushi suspects training now -- with new computer architecture -- would take less than a day, and cancer identification and delineation could be done in less than a minute per slide.
"To put this in perspective," Madabhushi said, "the machine could do the analysis during 'off hours,' possibly running the analysis during the night and providing the results ready for review by the pathologist when she/he were to come into the office in the morning."
Source-Eurekalert